-
自19世纪70年代,Rao[1]和Zafiriou[2]使用闪光光解在海水中检出卤素自由基以来,卤素自由基的检测技术及其环境化学过程引起研究者的普遍关注。据估计,表层海水中卤素自由基的总浓度约为2.0 × 10−14 mol·L−1,高于·OH浓度(1.1 × 10−17 mol·L−1)约3个数量级[3]。因此,卤素自由基在溶解性有机质(DOM)的卤化[4]、部分天然物质[5]和有机污染物[6-7]的降解过程中均可发挥重要作用。近年来,UV/氯高级氧化技术的发展,也使得氯自由基参与的有机污染物降解成为环境领域的研究热点问题之一[8-10]。卤素自由基参与的反应可导致卤代污染物的生成,而卤代化合物的环境持久性和毒性往往高于其对应的非卤代化合物[11-13]。因此,研究氯自由基和有机化合物的反应过程具有重要的环境意义。
氯原子(也称氯自由基,Cl·)和二氯阴离子自由基(Cl2·−)是具有较高氧化活性的卤素自由基,其标准氧化电势分别为2.47 V和2.00 V[14]。高的还原电势使得Cl·和有机化合物有着较高的反应活性,反应速率常数范围为108—1010 L·mol−1·s−1 [15-20]。Cl2·−和有机化合物的反应速率常数范围为< 103—1010 L·mol−1·s−1 [21-25]。卤素自由基与有机污染物的反应速率可影响天然水中污染物的环境持久性及UV/氯水处理过程中污染物的去除效率,获取卤素自由基与有机化学品的反应速率常数对评估其在污染物降解过程中的贡献至关重要。定量结构活性关系(QSAR)模型已经成为一种高效获取有机化合物各类反应速率常数的工具[26]。Lei等[9]构建了部分特定结构痕量有机污染物的Hammett常数与Cl·和Cl2·−反应速率常数的关系模型。然而,对于有机化合物与氯自由基反应速率常数的QSAR模型尚缺少系统的研究,氯自由基反应速率常数的数据仍很缺乏。
因此,本研究从文献中搜集了有机化合物分别与Cl·和Cl2·−的二级反应速率常数(
$ {\text{k}}_{\text{Cl·}} $ 和$ \text{}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ ),通过理论计算获取了有机化合物的分子结构描述符,采用多元线性回归(MLR)的方法构建了可预测Cl·和Cl2·−与有机物的反应速率常数的QSAR模型,通过Euclidean距离法以及Williams图法分别表征了模型的描述符域和应用域,并对模型进行了机理解释。 -
从文献[3,18,21,27,28]中收集了28种和18种有机污染物分别与Cl· (表1)和Cl2·−(表2)的二级反应速率常数。所有有机物的反应条件(pH=7、温度20 ℃)相同。如果不同的文献中有多个k值,则取平均值。根据文献[29],将数据随机分为训练集和验证集,两者的比例约为4:1。在分析之前,将k进行对数变换,以缩小取值范围,增加其匀称性。
-
在Gaussian 16[30]软件上采用B3LYP[24,26,29,31]泛函6-31 + G (d, p)基组对所选取的有机污染物进行分子结构优化,得到优化好的分子构型之后,计算量子化学描述符(如最高占据分子轨道能EHOMO,最低未占据分子轨道能ELUMO,偶极矩μ,极化率α等)。基于Gaussian软件优化好的构型,使用Dragon 7.0[32]软件计算表征分子结构信息的Dragon描述符。对所提取的Dragon描述符进行筛选,剔除常数项和近似项,以及缺失的描述符等不符合要求的描述符。
-
运用SPSS 19.0软件采用多元线性回归的方法筛选描述符,构建QSAR模型。最佳的模型需要满足以下几个条件[33]:1)分子结构描述符数量较少(训练集中的化合物数量/描述符的数量大于5);2)较高的决定系数(R2),一般大于0.6;3)方差膨胀因子(VIF)小于10,VIF表示预测变量的多重共线性;4)显著性水平(P)小于0.05。
对所建模型采用内部验证和外部验证来评价其相关性能。对于内部验证来说,相关的统计学参数要满足以下几个原则:内部交叉验证系数(Q2LOO)和Q2BOOT(由bootstrapping计算得到的参数)大于0.6,较高的校正后的决定系数(R2adj),一般大于0.7,较小的均方根误差(RMSE)。对于外部验证来说,具有较高的验证集外部可解释方差Q2ext以及验证集决定系数R2ext,较小的均方根误差(RMSE)。相关统计学参数的计算公式如下:
其中,n是样本数;p是自变量的个数;
$ \widehat{{y}_{\mathrm{i}}} $ 和yi分别是化合物活性的预测值和实测值;$ \stackrel{-}{{y}_{i}} $ 是化合物活性实测值的平均值。模型的应用域(ADs)是使模型可靠和可预测的化学结构空间,本研究用Euclidean距离法对模型的描述符域进行了表征,其计算公式如下:
其中,μ为描述符x的均值。
本模型的应用域采用基于Williams方法进行表征,杠杆指及其警戒值的的公式定义如下:
其中,n为训练集化合物数量,k为模型中的描述符的个数,Xi为第i个有机物分子结构描述符值。
本研究最大警戒值h*,取|δ| > 3为离群点。
-
$ \text{lg}{\text{k}}_{\text{Cl·}} $ 的QSAR模型为:$ \text{lg}{\text{k}}_{\text{Cl·}}\text{=} $ 11.133 - 7.236 × Dv - 0.087 × VE3sign_RG + 0.569 × GATS5e - 0.158 × H -051ntr = 22, R2adj, tr = 0.810, Q2LOO = 0.716, Q2BOOT = 0.653, RMSEtr = 0.34, Kx = 0.127, Kxy = 0.303; next = 6, R2adj, ext = 0.861, RMSEext = 0.48, Q2ext = 0.793
$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ 的QSAR模型为:$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ = 3.657 + 0.374 × RDF010s - 0.037 × ATSC5s + 0.073 × L/Bwntr = 14, R2adj, tr = 0.902, Q2LOO = 0.839, Q2BOOT = 0.735, RMSEtr = 0.29, Kx = 0.385, Kxy = 0.401; next = 4, R2adj, ext = 0.838, RMSEext = 0.67, Q2ext = 0.629
关于
$ \text{lg}{\text{k}}_{\text{Cl·}} $ QSAR模型,模型包含4个分子结构描述符,分别是Dv、VE3sign_RG、GATS5e和H-051,所有的分子结构描述符都属于Dragon描述符。其中ntr和next分别表示训练集和验证集中有机化合物的数量。对于模型的内部验证,R2adj, tr > 0.60表示训练集具有高拟合优度。交叉验证系数(Q2LOO)和Q2BOOT值均大于0.6,表明模型具有良好的统计稳健性。由表1可知VIF < 10并且Kxy > Kx,因此所建立的模型不存在多重共线性。$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ QSAR模型同$ \text{lg}{\text{k}}_{\text{Cl·}} $ ,所有的统计学参数都满足条件,模型具有较高的拟合优度,稳健性以及预测能力。如图1和图2所示,模型预测值与文献实验数据吻合较好。因此,所建立的模型符合经济合作与发展组织(OECD)原则的要求,能够准确地预测有机化合物与Cl·与Cl2·−反应的速率常数。 -
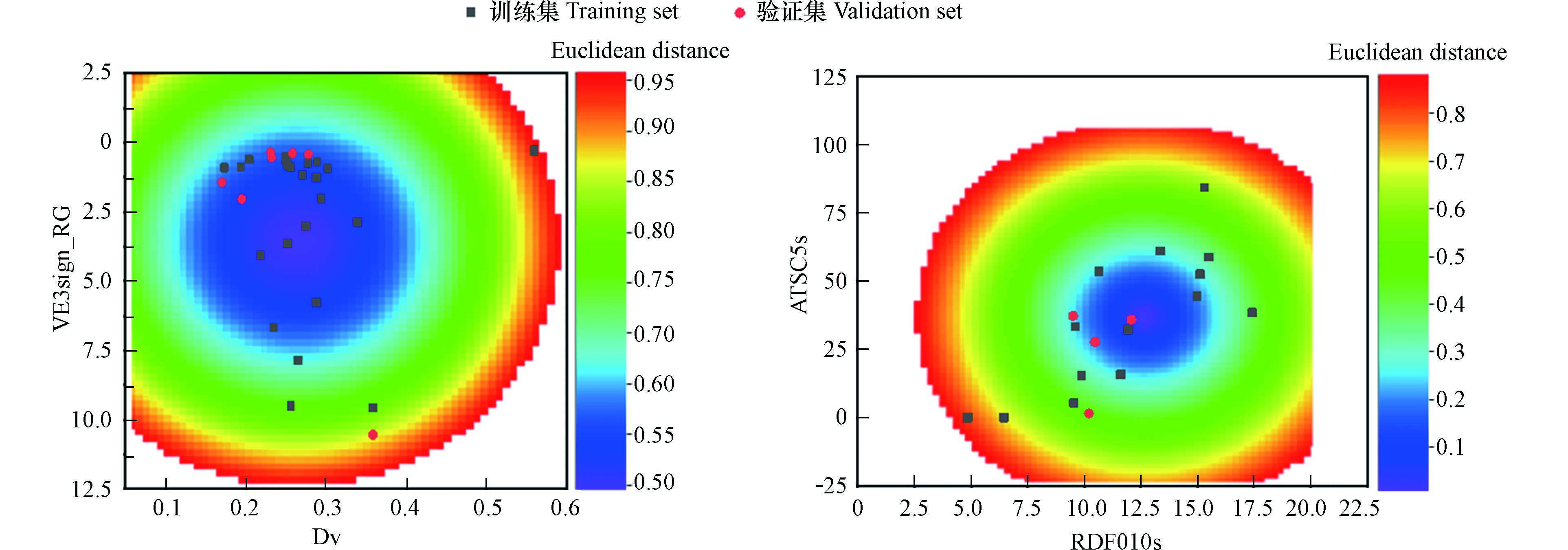
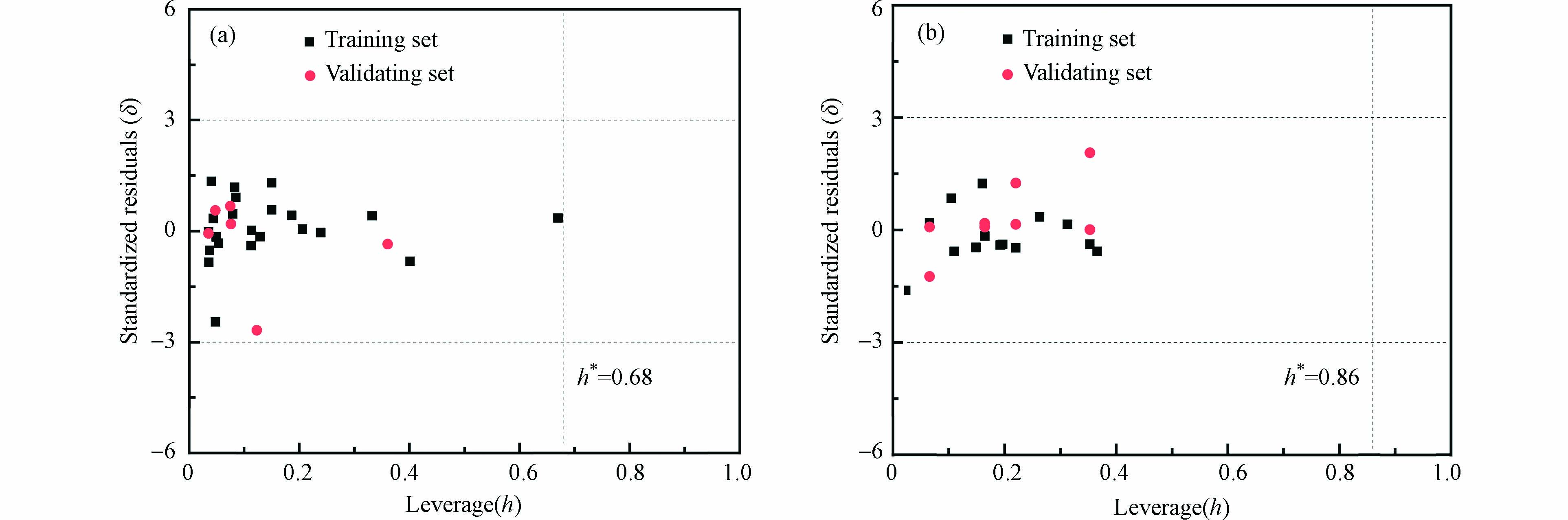
采用基于欧几里得距离表征的描述符域以及Williams的方法对模型的应用域进行表征,如图2和图3所示,两个模型所有的有机化合物都在描述符内,并且在描述符域的平面空间内分布比较均匀。
$ \text{lg}{\text{k}}_{\text{Cl·}} $ 模型中,所有化合物均在在警戒值范围h* = 0.68之内,且|δ| < 3。因此所构建的模型能预测含有羟基、羧基、羰基以及芳香族化合物与Cl·的二级反应速率常数。而在$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ 模型中,所有的化合物也都处于应用域内,由于本次建模所选取的化合物大部分为苯甲酸类物质,因此所构建的QSAR模型可以较好地预测苯甲酸类物质与Cl2·−的二级反应速率常数。 -
由表1可知,所搜集的Cl·与有机物反应的速率常数范围为9.3 × 106—4 × 1010 L·mol−1·s−1。芳香族化合物的速率常数要大于小分子的烯烃、醚、羧酸、醇类化合物,并且醇类有机物与Cl·的反应速率常数要明显高于小分子的烯烃、醚类、羧酸类有机物。由表2可知,Cl2·−与表中有机物的反应速率常数范围为1.9 × 106—1.1 × 109 L·mol−1·s−1,由表可知对氯苯甲酸要小于对溴苯甲酸,含氨基的苯甲酸类物质(1.1 × 109 L·mol−1·s−1)要高于含羟基(2.8 × 108 L·mol−1·s−1)和甲氧基类(2.0 × 108 L·mol−1·s−1)物质。对比表1与表2中的反应速率常数可以得出,有机物与Cl·的反应速率常数整体上要大于Cl2·−的反应速率常数,对于两表中含有的共同化合物来说,与Cl·的反应速率常数要远远大于与Cl2·−的反应速率常数,是由Cl·和Cl2·−的氧化还原电势所决定的[14]。
从表3可以看出,对于
$ \text{lg}{\text{k}}_{\text{Cl·}} $ 模型,描述符Dv的t值(t = -13.517)最大,表明描述符Dv在这个模型中有最重要的作用。Dv描述符属于WHIM (weighted holistic invariant molecular)描述符[34-35],这种描述符的建立是为了获取有关分子的三维信息,包括分子的大小、形状、对称性和原子分布。在本次研究中,Dv定义为D总可达性指数/原子范德华体积加权(D total accessibility index / weighted by atomic van der Waals volumes),此描述符前面的系数为负表明化合物的Dv值越大,化合物与氯自由基的反应速率常数越小。VE3sign_RG定义为来自倒数平方几何矩阵的最后一个特征向量的对数系数和(logarithmic coefficient sum of the last eigenvector from reciprocal squared geometrical matrix),此描述符属于3D matrix-based描述符,系数为负数表明此描述符与反应速率常数呈负相关;GATS5e (Geary autocorrelation—lag 5 / weighted by atomic Sanderson electronegativities) [35-38]为Sanderson电负性加权的lag5 Geary 自相关指数,属于二维自相关描述符,在这个描述符中,Geary系数是一个距离型函数,该函数是分子中每个原子计算的任何物理化学性质,如原子质量、极化率等。因此,分子原子代表了空间中离散点的集合和在这些点上的函数的原子性质,描述符前面的系数为正,代表其值越高的分子在结构中电子分布越合适,而已知Cl·较强的电子受体能力可以使难降解的化合物降解,因此其值与反应速率常数呈正相关。H-051 (H attached to alpha-C Atom-centred fragments)为H加到以原子为中心的alpha-c片段的基本描述符,系数为负,代表化合物alpha-c上的H原子的数量与反应速率常数呈负相关,如苯酚、苯胺、六甲基苯、苯甲酸的alpha-c数量的氢原子较二氯甲烷,甲烷二醇等结构的少,因此速率常数较大。从表4可以看出,对于
$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ 模型来说,描述符RDF010s (radial distribution function weighted 010s / weighted by 1-state)的t值(t = 8.311)最大,是影响本模型最大的描述符,此描述符代表I-stat加权的径向分布描述符[39],与反应速率常数呈负相关。剩余两个描述符分别为ATSC5s和L/Bw,其意义分别为I-state加权的lag5中心[40-42]Broto-Moreau自相关描述符以及用几何描述符的形状指数来计算长宽比[43-44],分别与反应速率常数呈负相关与正相关。 -
综上可知,本研究采用MLR方法,发展了一种在OECD原则下实现快速且便捷地预测污染物与卤素自由基反应速率常数的QSAR模型,并通过Euclidean距离法以及Williams方法表征了模型的描述符域及应用域。本模型的的拟合优度较高,稳健性以及预测性都比较好,能够较为准确地预测醇类、醚类、醛类、酸类烷烃及芳香族化合物室温条件下的
$ {\text{k}}_{\text{Cl·}} $ 值以及苯甲酸取代类物质的$ {\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ 值。本研究可为进一步发展优化基于卤素自由基的有机污染物去除技术如高级氧化技术提供科学依据,为评估卤素自由基引发的表层海水中有机污染物毒理性变化及生态风险变化提供基础数据。
水相中有机化学品和卤素自由基反应速率常数的定量结构-活性关系模型
A quantitative structure-activity relationship model for reaction rate constants of organic chemicals with halogen radicals in aqueous phase
-
摘要: 卤素自由基尤其是氯自由基在污水中难降解污染物的降解以及表层海水中环境微污染物的降解过程中发挥重要作用,能够影响污染物的环境持久性与毒性风险。在污水及表层海水中存在着多种多样的有机污染物,单纯依靠实验手段测定它们和氯自由基的反应速率常数费时费力并且成本较高。因此本研究通过发展氯自由基与有机污染物的反应速率常数的QSAR模型来实现反应速率常数的定量预测。本研究搜集了28种有机化合物与Cl·的反应速率常数以及18种有机化合物与Cl2·−的反应速率常数的实测值,采用多元线性回归(MLR)分别建立了QSAR模型。
$ \text{lg}{\text{k}}_{\text{Cl·}}= $ 11.133 − 7.236 × Dv − 0.087 × VE3sign_RG + 0.569 × GATS5e − 0.158 × H-051。$ \text{lg}{\text{k}}_{\text{Cl·}} $ 的QSAR模型的校正决定系数(R2adj, tr)为0.810,均方根误差(RMSEtrain)为0.34,去一法交叉验证系数(Q2LOO)为0.716,外部验证系数Q2 ext = 0.793;$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ = 3.657 + 0.374 × RDF010s − 0.037 × ATSC5s + 0.073 × L/Bw。$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ QSAR模型R2adj, tr为0.902,RMSEtrain为0.29,Q2LOO为0.839,Q2ext为0.629。表明两个模型具有较高的拟合优度、稳健性和预测能力。应用域表征结果显示所选择的模型具有较好的代表性,所有化合物均在应用域范围内。所构建的模型可以实现含羟基,羰基,羧基等多种官能团有机化合物$ {\text{k}}_{\text{Cl·}} $ 及$ {\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ 的定量预测。-
关键词:
- 氯自由基 /
- 反应速率常数 /
- 多元线性回归 /
- 定量结构活性关系模型
Abstract: Halogen radicals, especially chlorine radicals, play an important role in the transformation and degradation of micro-pollutants in sewage and surface seawater, which can affect the persistence and toxicity risk of pollutants in water environment. There are many kinds of organic pollutants in aqueous phase. However, it is time-consuming and costly to determine the reaction rate constants of organic pollutants with chlorine radicals based on the experiments. Quantitative structure-activity relationship (QSAR) is well recognized as a valuable tool to achieve rapidly quantitative prediction of these reaction rate constants. In this study, the reaction rate constants of 28 organic compounds with Cl· and 18 organic compounds with Cl2·− were collected. QSAR models were then established by multiple linear regression (MLR) for these compounds, respectively (i.e.$ \text{lg}{\text{k}}_{\text{Cl·}}= $ 11.133 − 7.236 Dv − 0.087 VE3sign_RG + 0.569 GATS5e − 0.158 H-051 and$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ = 3.657 + 0.374 RDF010s − 0.037 ATSC5s + 0.073 L/Bw). It shows that these two models have good fit, robustness and strong prediction ability, with the adjusted determination coefficients (R2adj, tr) of 0.810 and 0.902, the root-mean-square errors (RMSEtrain) of 0.34 and 0.29, leave-one-out cross validated coefficients (Q2LOO) of 0.716 and 0.839, and the external validation coefficients (Q2ext) of 0.793 and 0.629, respectively. The applicability domain of the two models reveals that the developed models have better representation, and all of compounds are in the applicability domain. The models constructed in this paper can be used in the quantitative prediction of$ {\text{k}}_{\text{Cl·}} $ and$ {\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}}\text{} $ reaction rate constants for many types of organic chemicals, such as compounds with functional groups of hydroxyl, carbonyl and carboxyl. -
-
图 2 基于欧几里得距离表征的
$ \text{lg}{\text{k}}_{\text{Cl·}} $ 和$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ 模型的描述符域Figure 2. Descriptor domain of the developed
$ \text{lg}{\text{k}}_{\text{Cl·}} $ and$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ QSAR model described by Euclidean distance-based method表 1 有机化合物的CAS号、名称、结构及与Cl•的二级反应速率常数(
$ k_{\text{Cl·}} $ Table 1. CAS, name, structure and rate constant of organic compounds with Cl•
序号
NumberCAS号 物质名称
Name结构
Structure$k_{\text{Cl·} }$
(L·mol−1·s−1)1 75-09-2 二氯甲烷 
9.3 × 106 [18] 2 67-64-1 丙酮* 
7.8 × 107 [18] 3 64-19-7 乙酸* 
1.0 × 108 [18] 4 78-93-3 2-丁酮 
2.4 × 108 [18] 5 79-09-4 丙酸 
1.2 × 109 [18] 6 60-29-7 乙醚 
1.3 × 109 [18] 7 1634-04-4 甲基叔丁基醚 
1.3 × 109 [18] 8 463-57-0 甲烷二醇 
1.4 × 109 [18] 9 14277-97-5 软骨藻酸 
1.5 × 109 [3] 10 75-65-0 叔丁醇 
1.5 × 109 [18] 11 79-31-2 异丁酸 
1.7 × 109 [18] 12 71-23-8 正丙醇 
2.2 × 109 [18] 13 64-17-5 乙醇 
2.2 × 109 [18] 14 64-18-6 甲酸 
2.8 × 109 [18] 15 67-63-0 异丙醇* 
3.7 × 109 [3] 16 617-86-7 三乙基硅烷 
3.8 × 109 [28] 17 78-92-2 仲丁醇 
5.0 × 109 [18] 18 150-76-5 对羟基苯甲醚 
5 × 109 [28] 19 67-56-1 甲醇* 
5.7 × 109 [18] 20 67-68-5 二甲基亚砜 
7 × 109 [28] 21 71-41-0 1-戊醇 
7.9 × 109 [28] 22 110-82-7 环己烷 
8 × 109 [28] 23 110-83-8 环己烯 
1.2 × 1010 [28] 24 65-85-0 苯甲酸* 
1.8 × 1010 [27] 25 151-10-0 间苯二甲醚 
2 × 1010 [28] 26 87-85-4 六甲基苯 
2.2 × 1010 [28] 27 108-95-2 苯酚* 
2.5 × 1010 [3] 28 62-53-3 苯胺 
4 × 1010 [28] 注:*为验证集化合物,其余为训练集化合物.
*Represents that the compounds are in the validating set. The other compounds are in the training set.表 2 有机化合物的CAS号、名称、结构及与Cl2•−的二级反应速率常数
Table 2. CAS, name, structure and rate constant of organic compounds with Cl2•−
序号
NumberCAS 物质名称
Name物质结构
Structure$k_{ {\text{Cl} }_{\text{2} }{\text{·} }^{\text{-} } }$
(L·mol−1·s−1)1 64-18-6 甲酸 
1.9 × 106 [21] 2 65-85-0 苯甲酸* 
2.0 × 106 [21] 3 107-13-1 丙烯腈 
2.2 × 106 [21] 4 74-11-3 对氯苯甲酸* 
3 × 106 [21] 5 110-17-8 富马酸 
4 × 106 [21] 6 99-94-5 对甲基苯甲酸 
5 × 106 [21] 7 619-65-8 对氰基苯甲酸 
5 × 106 [21] 8 100-21-0 对苯二甲酸 
6 × 106 [21] 9 586-76-5 4-溴苯甲酸 
7 × 106 [21] 10 79-10-7 丙烯酸 
1.9 × 107 [21] 11 103-84-4 N-乙酰苯胺 
2.0 × 107 [21] 12 4436-74-2 反式-2-丁烯-1,4-二甲酸 
1.6 × 108 [21] 13 100-09-4 对甲氧基苯甲酸* 
2.0 × 108 [21] 14 3588-17-8 反式,反式-1,3-丁二烯-1,4-二羧酸 
2.1 × 108 [21] 15 99-96-7 对羟基苯甲酸 
2.8 × 108 [21] 16 107-18-6 丙烯醇* 
5.9 × 108 [21] 17 110-44-1 山梨酸 
6.8 × 108 [21] 18 150-13-0 对氨基苯甲酸 
1.1 × 109 [21] 注:*为验证集化合物,其余为训练集化合物.
*Represents that the compounds are in the validating set. The other compounds are in the training set.表 3
$ \text{lg}{\text{k}}_{\text{Cl·}} $ Table 3. Meaning of descriptors, coefficients, VIF values, t values and p in the
$ \text{lg}{\text{k}}_{\text{Cl·}} $ 描述符
Descriptors描述符意义
Descriptor meaning系数
CoefficientVIF t p Dv 与分子范德华体积相关的3D结构描述符 −7.236 1.022 −13.517 ﹤1×10−3 VE3sign_RG 来自倒数平方几何矩阵的最后一个特征向量的对数系数和 −0.087 1.095 −5.630 ﹤1×10−3 GATS5e Sanderson电负性加权的lag5 Geary 自相关指数 0.569 1.068 5.719 ﹤1×10−3 H-051 H加到以原子为中心的alpha-c片段的基本描述符 −0.158 1.032 −4.966 ﹤1×10−3 表 4
$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}} $ Table 4. Meaning of descriptors, coefficients, VIF values, t values and p in the
$ \text{lg}{\text{k}}_{{\text{Cl}}_{\text{2}}{\text{·}}^{\text{-}}}\text{} $ 描述符
Descriptors描述符意义
Descriptor meaning系数
CoefficientVIF t p RDF010s 0.10I-stat加权的径向分布描述符 0.374 2.484 8.311 ﹤1×10−3 ATSC5s I-state加权的lag5中心Broto-Moreau自相关描述符 −0.037 2.548 −4.885 ﹤1×10−3 L/Bw 用几何描述符的形状指数来计算长宽比 0.073 1.051 3.064 ﹤1×10−3 -
[1] RAO P. Flash photolysis of seawater [J]. Current Science, 1973, 42(17): 599. [2] ZAFIRIOU O C. Sources and reactions of OH and daughter radicals in seawater [J]. Journal of Geophysical Research, 1974, 79(30): 4491-4497. doi: 10.1029/JC079i030p04491 [3] PARKER K M, MITCH W A. Halogen radicals contribute to photooxidation in coastal and estuarine waters [J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(21): 5868-5873. doi: 10.1073/pnas.1602595113 [4] JOE-WONG C, SCHLESINGER D R, CHOW A T, et al. Sea level rise produces abundant organobromines in salt-affected coastal wetlands [J]. Geochemical Perspectives Letters, 2019, 10: 31-35. [5] PARKER K M, REICHWALDT E S, GHADOUANI A, et al. Halogen radicals promote the photodegradation of microcystins in estuarine systems [J]. Environmental Science & Technology, 2016, 50(16): 8505-8513. [6] JIN R, ZHENG M, LAMMEL G, et al. Chlorinated and brominated polycyclic aromatic hydrocarbons: Sources, formation mechanisms, and occurrence in the environment [J]. Progress in Energy and Combustion Science, 2020, 76: 100803. doi: 10.1016/j.pecs.2019.100803 [7] ZHAO Q, FANG Q, LIU H, et al. Halide-specific enhancement of photodegradation for sulfadiazine in estuarine waters: Roles of halogen radicals and main water constituents [J]. Water Research, 2019, 160: 209-216. doi: 10.1016/j.watres.2019.05.061 [8] GUO K, WU Z, SHANG C, et al. Radical chemistry and structural relationships of PPCP degradation by UV/chlorine treatment in simulated drinking water [J]. Environmental Science & Technology, 2017, 51(18): 10431-10439. [9] LEI Y, CHENG S, LUO N, et al. Rate constants and mechanisms of the reactions of Cl• and Cl2•− with trace organic contaminants [J]. Environmental Science & Technology, 2019, 53(19): 11170-11182. [10] ZHANG R, MENG T, HUANG C H, et al. PPCP degradation by chlorine-UV processes in ammoniacal water: New reaction insights, kinetic modeling, and DBP formation [J]. Environmental Science & Technology, 2018, 52(14): 7833-7841. [11] JIANG J, HAN J, ZHANG X. Nonhalogenated aromatic DBPs in drinking water chlorination: A gap between NOM and halogenated aromatic DBPs [J]. Environmental Science & Technology, 2020, 54(3): 1646-1656. [12] 耿利鸣, 马广才, 尉小旋, 等. 卤代有机污染物抑制甲状腺激素代谢酶活性的研究进展 [J]. 生态毒理学报, 2019, 14(4): 14-22. GENG L M, MA G C, WEI X X, et al. Inhibition of metabolic activities of thyroid hormones by halogenated organic contaminants [J]. Asian Journal of Ecotoxicology, 2019, 14(4): 14-22(in Chinese).
[13] 林坤德, 陈艳秋, 袁东星. 新型污染物卤代咔唑的环境行为及生态毒理效应 [J]. 环境科学, 2016, 37(4): 1576-1583. LIN D K, CHEN Y Q, YUAN D X. Environmental behaviors and ecotoxicology of the emerging contaminants polyhalogenated carbazoles [J]. Environmental Science, 2016, 37(4): 1576-1583(in Chinese).
[14] XIANG Y Y, FANG J Y, SHANG C. Kinetics and pathways of ibuprofen degradation by the UV/chlorine advanced oxidation process [J]. Water Research, 2016, 90: 301-308. doi: 10.1016/j.watres.2015.11.069 [15] LERI A C, MAYER L M, THORNTON K R, et al. A marine sink for chlorine in natural organic matter [J]. Nature Geoscience, 2015, 8: 620-624. doi: 10.1038/ngeo2481 [16] BUXTON G V, BYDDER M, SALMON G A, et al. The reactivity of chlorine atoms in aqueous solution. Part III. The reactions of Cl· with solutes [J]. Physical Chemistry Chemical Physics, 2000, 2(2): 237-245. doi: 10.1039/a907133d [17] BUXTON G V, WANG J, SALMON G A. Rate constants for the reactions of NO3·-, SO4·- and Cl· radicals with formate and acetate esters in aqueous solution [J]. Physical Chemistry Chemical Physics, 2001, 3(13): 2618-2621. doi: 10.1039/b101932p [18] WICKTOR F, DONATI A, HERRMANN H, et al. Laser-based spectroscopic and kinetic investigations of reactions of the Cl atom with oxygenated hydrocarbons in aqueous solution [J]. Physical Chemistry Chemical Physics, 2003, 5(12): 2562-2572. doi: 10.1039/b212666d [19] ALEGRE M L, GERONES M, ROSSO J A, et al. Kinetic study of the reactions of chlorine atoms and Cl2·- radical anions in aqueous solutions. 1. reaction with benzene [J]. The Journal of Physical Chemistry A, 2000, 104(14): 3117-3125. doi: 10.1021/jp9929768 [20] MARTIRE D O, ROSSO J A, BERTOLOTTI S, et al. Kinetic study of the reactions of chlorine atoms and Cl2·- radical anions in aqueous solutions. II. toluene, benzoic acid, and chlorobenzene [J]. The Journal of Physical Chemistry A, 2001, 105(22): 5385-5392. doi: 10.1021/jp004630z [21] HASEGAWA K, NETA P. Rate constants and mechanisms of reaction of Cl2·- radicals [J]. The Journal of Physical Chemistry, 1978, 82(8): 854-857. doi: 10.1021/j100497a003 [22] ADAMS G E, ALDRICH J E, BISBY R H, et al. Selective free radical reactions with proteins and enzymes: reactions of inorganic radical anions with amino acids [J]. Radiation Research, 1972, 49(2): 278-289. doi: 10.2307/3573266 [23] NETA P, HUIE R E, ROSS A B. Rate constants for reactions of inorganic radicals in aqueous solution [J]. Journal of Physical and Chemical Reference Data, 1988, 17(3): 1027-1284. doi: 10.1063/1.555808 [24] MERENYI G, LIND J. Reaction mechanism of hydrogen abstraction by the bromine atom in water [J]. Journal of the American Chemistry Society, 1994, 116(17): 7872-7876. doi: 10.1021/ja00096a050 [25] ALFASSI Z B, HUIE R E, MARGUET S, et al. Rate constants for reactions of iodine atoms in solution [J]. International Journal of Chemistry Kinetics, 1995, 27(2): 181-188. doi: 10.1002/kin.550270208 [26] 郑珊珊, 李田田, 王晶, 等. 芳香族化合物与水合电子水相反应速率常数的QSAR模型研究 [J]. 环境化学, 2019, 38(5): 1005-1013. ZHENG S S, LI T T, WANG J, et al. QSAR models for predicting the aqueous reaction rate constants of aromatic with hydrated electrons [J]. Environmental Chemistry, 2019, 38(5): 1005-1013(in Chinese).
[27] FANJ J Y, FU Y, SHANG C. The roles of reactive species in micropollutant degradation in the UV/free chlorine system [J]. Environmental Science & Technology, 2014, 48(3): 1859-1868. [28] ALFASSI Z B, MOSSERI S, NETA P. Reactivities of chlorine atoms and peroxyl radical formed in the radiolysis of dichloromethane [J]. The Journal of Physical Chemistry, 1989, 93(4): 1380-1385. doi: 10.1021/j100341a040 [29] LI C, ZHENG S S, LI T T, et al. Quantitative structure-activity relationship models for predicting reaction rate constants of organic contaminants with hydrated electrons and their mechanistic pathways [J]. Water Research, 2019, 151: 468-477. doi: 10.1016/j.watres.2018.12.010 [30] FRISCH M J, TRUCKS G W, SEHLEGEL H B, et a1. Gaussian 09, revision a. 02[CP] . Wallingford, CT: Gaussian, Inc, 2009. [31] ZHANG Y, WANG J, CHEN J, et al. Phototransformation of 2, 3-dibromopropyl-2, 4, 6-tribromophenyl ether (DPTE) in natural waters: Important roles of dissolved organic matter and chloride ion [J]. Environmental Science & Technology, 2018, 52(18): 10490-10499. [32] SOPER-HOPPER M T, PETROV A S, HOWARD J N, et al. Collision cross section predictions using 2-dimensional molecular descriptors [J]. Chemical Communications, 2017, 53: 7624-7627. doi: 10.1039/C7CC04257D [33] XIAO R, YE T, WEI Z, et al. Quantitative structure−activity relationship (QSAR) for the oxidation of trace organic contaminants by sulfate radical [J]. Environmental Science & Technology, 2015, 49: 13394-13402. [34] TODESCHINI R, GRAMATICA P. 3D-modelling and Prediction by WHIM Descriptors. Part 6. Application of WHIM Descriptors in QSAR Studies [J]. Molecular Informatics, 1997, 16(2): 107-192. [35] YANG C, SHAO Y H, ZHI X Y, et al. Semisynthesis and quantitative structure-activity relationship (QSAR) study of some cholesterol-based hydrazone derivatives as insecticidal agents [J]. Bioorganic & Medicinal Chemistry Letters, 2013, 23(17): 4806-4812. [36] YANGJEH A H. QSAR study of the 5-HT1A receptor affinities of aryl piperazines using a genetic algorithm–artificial neural network model [J]. Monatshefte für Chemie Chemical Monthly, 2009, 140(5): 523-530. [37] JOSE L. VELAZQUEZ-L J C, et al. Estimation of 2D autocorrelation descriptors and 2D Monte Carlo descriptors as a tool to build up predictive models for acetylcholinesterase (AChE) inhibitory activity [J]. Chemometrics and Intelligent Laboratory Systems, 2019, 184: 14-21. doi: 10.1016/j.chemolab.2018.11.008 [38] GOKHALE N, JAIN S. QSAR studies on thiosemicarbazone derivatives as an anticancer agent [J]. International Journal of Pharmacy Teaching & Practices, 2015, 6(2): 2111-2119. [39] FATEMI M H, DOROSTKAR F. QSAR prediction of D2 receptor antagonistic activity of 6-methoxy benzamides [J]. European Journal of Medicinal Chemistry, 2010, 45(11): 4856-4862. doi: 10.1016/j.ejmech.2010.07.056 [40] USMAN B, MAAROF H, ABDALLAH H H, et al. Computational evaluation of the effect of structural parameters of 3-flouro thiophene and 3-thiophene malonic acid on corrosion inhibition efficiency of mild steel in acidic media [J]. International Journal of Electrochemical Science, 2015, 10: 3223-3229. [41] BORONOVÁA K, LEHOTAY J, HROBONOVÁ K, et al. Study of physicochemical interaction of aryloxyaminopropanol derivatives with teicoplanin and vancomycin phases in view of quantitative structure-property relationship studies [J]. Journal of Chromatography A, 2013, 1301: 38-47. doi: 10.1016/j.chroma.2013.05.046 [42] SAAVEDRA L M, ROMANELLI G P, ROZO C E, et al. The quantitative structure–insecticidal activity relationships from plant derived compounds against chikungunya and zika Aedes aegypti (Diptera: Culicidae) vector [J]. Science of the Total Environment, 2018, 610-611: 937-943. doi: 10.1016/j.scitotenv.2017.08.119 [43] PEREIRA F, LATINO D A R S, AIRES-DE-SOUSA J. Estimation of mayr electrophilicity with a quantitative structure−property relationship approach using empirical and dft descriptors [J]. The Journal of Organic Chemistry, 2011, 76(22): 9312-9319. doi: 10.1021/jo201562f [44] GONZÁlLEZ P M, GÁNDARA Z, FALL Y, et al. Radial distribution function descriptors for predicting affinity for vitamin D receptor [J]. European Journal of Medicinal Chemistry, 2008, 43(7): 1360-1365. doi: 10.1016/j.ejmech.2007.10.020 -

















 下载:
下载: